The first paper from my PhD, titled “Confirmation bias through selective overweighting of choice-consistent evidence” is just out in Current Biology. A blogpost explaining the paper and its findings, co-written by me, can be found here. In the blogpost below, I share the methodological insights I gained while working on the project, and some mistakes that otherwise hindered my progress. The purpose of the blog is to inform early career researchers such as myself about some potential pitfalls, and efficient practices to get the best out of their time.
1. Model comparison using a single measure (for e.g. Bayes Information Criterion, BIC) need not always reveal the correct model for the data. In the paper, model comparison using BIC (Figure 1A) was further corroborated by model simulations. Specifically, we simulated data using the best fitting parameters (Figure 1B), and with a range of parameters (Figure 1C) in each model. We then investigated for the presence of the model-free ROC index, a diagnostic feature of behavioural data, in the simulated data (Figure 1B, C below). This allowed us to suggest with certainty that Choice-based Selective Gain model accounted for the data.
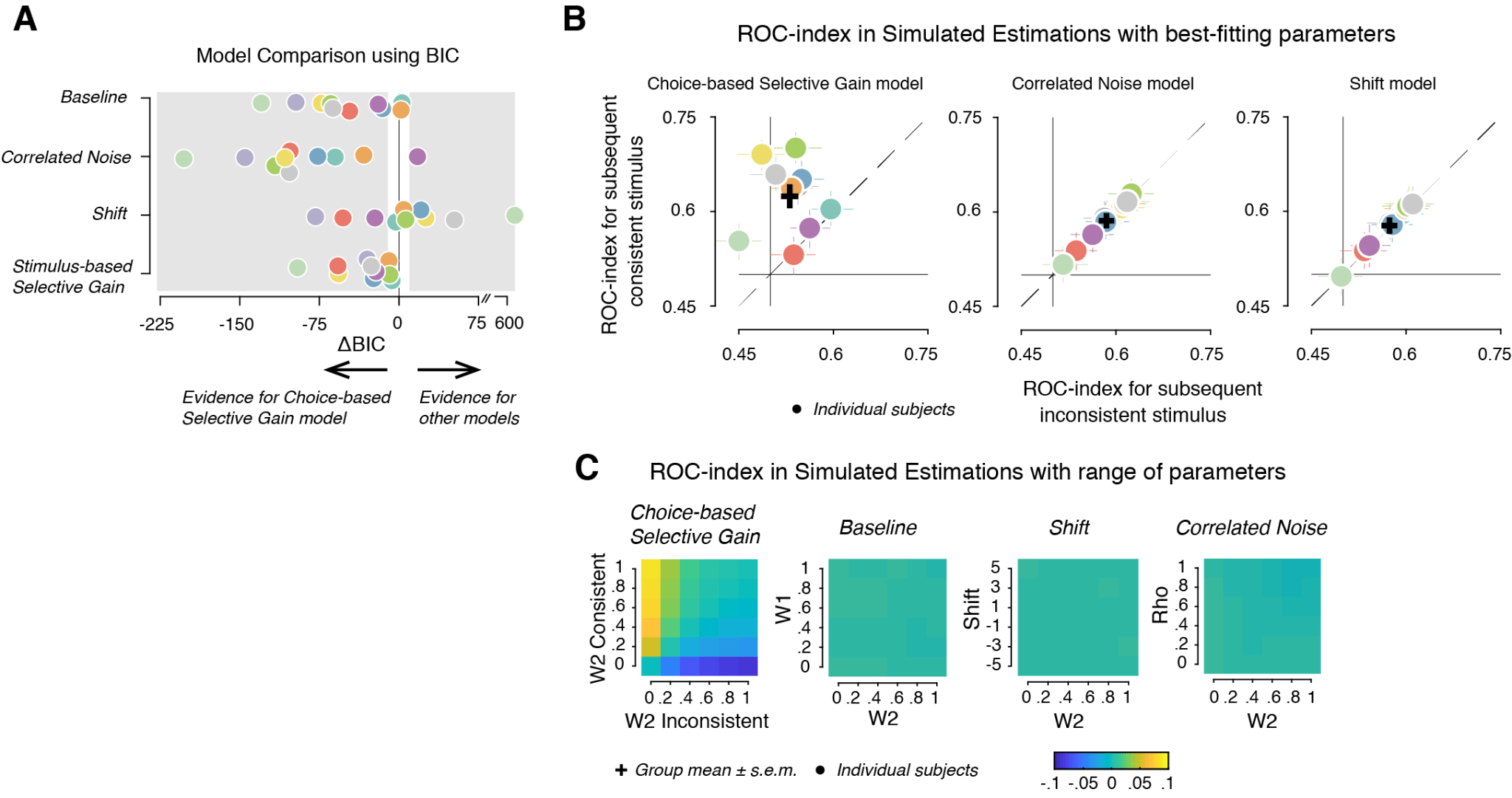
Figure 1
2. If and where possible, use non-parametric measures for statistical tests, and for model-free analyses. In the study, we had a continuous estimate as the behavioural measure in every trial. To compare how these estimates differ across two different conditions of interest, we used the ROC index (illustrated below, Figure 2A), instead of mean (or median) and standard deviation. ROC-index quantifies the separability of two distributions (Figure 2B). ROC curves (Figure 2A, right panel) were constructed by shifting a criterion across both distributions (Figure 2A, left panel), and plotting against one another, for each position of the criterion, the fraction of trials for which responses were larger than the criterion in each distribution. The area under the resulting ROC curve (AUC; grey shading), here referred to as ‘ROC-index’, quantifies the probability with which an ideal observer can predict the distribution from the single-trial estimation. An ROC-index of 0.5 implies that the distributions are inseparable (dashed line in Figure 2A, right panel). It is important to note that ROC indices between the conditions differ from 0.5 when either the means or the spread of the distributions are different. Two model-based measures (weights and noise, Figure 2C) explain the effect captured by the ROC-indices.
Figure 2

3. Simulated data can be used to validate model-free measures. This is a common practice by researchers developing methods, where simulated data with a null effect is used to validate a new method. We established ROC-index as a diagnostic feature of confirmation bias in our dataset after we validated it with data simulated using the Baseline model, which by design does not differentiate the two conditions (Figure 1C, second panel from the right).
4. An often overlooked aspect of modelling studies is the ability of models to recover parameters from simulated data. Specifically, the model should recover individual parameters faithfully (visualising the correlation between the actual and recovered parameters helps), and the recovered parameters should not exhibit correlations that were non-existent between the actual parameters. Parameter recovery is the reason we ruled out the Extended Choice-based Selective Gain model, the most complex model in the paper (see section Parameter Recovery in STAR methods in the paper), as the recovered parameters exhibited spurious correlations.
5. While doing a fixed-effects analyses (by treating the sample population as one subject), bootstrapping to obtain confidence intervals for the parameter estimates allowed us to better understand the difference between the estimates. On the same note, obtaining confidence intervals for the parameter estimates in each individual subject gave us confidence about the effect-size in each subject.
6. The last, but a very important aspect, of every project is adopting good coding practices. Specifically, in the project, I found it extremely useful to check every snippet of code to make sure it is doing what it is supposed to do, possibly by visualising the outputs, before starting to do computationally extensive analyses. I am currently working on an fMRI project and this practice saved a lot of time and effort allowing me to identify any bugs, and check if the pipeline introduces any unintended artefacts in the analyses.
Before I end, a note of thanks to Konstantinos Tsetsos, and Anne Urai, my code-buddies, who taught me some of the above points.